Research
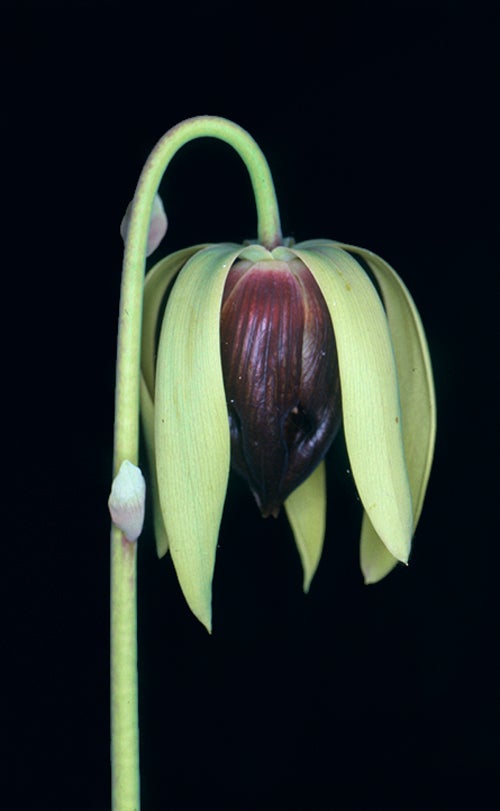
We study evolutionary processes in the crucible of nature, always with an eye towards applications to real-world problems. Variation within and between populations over geographic latitudes from Florida to northern Manitoba  and from sea level to the Appalachian highlands provides abundant end-products of natural evolutionary events. Questions asked in our laboratory are encompassed in the broader question, “What is the molecular genetic basis of evolutionary change in natural populations?” Our primary research organism is Wyeomyia smithii, a small mosquito that develops only within the water-filled leaves of the purple pitcher plant. Facilities unique to our lab include computer-controlled environment rooms unlike any others in the world, in which we are able to recreate any environment from the tropics to the polar regions of Earth. We use a variety of quantitative genetic, molecular, and genomic tools to address a broad spectrum of questions including the following ongoing research projects:
and from sea level to the Appalachian highlands provides abundant end-products of natural evolutionary events. Questions asked in our laboratory are encompassed in the broader question, “What is the molecular genetic basis of evolutionary change in natural populations?” Our primary research organism is Wyeomyia smithii, a small mosquito that develops only within the water-filled leaves of the purple pitcher plant. Facilities unique to our lab include computer-controlled environment rooms unlike any others in the world, in which we are able to recreate any environment from the tropics to the polar regions of Earth. We use a variety of quantitative genetic, molecular, and genomic tools to address a broad spectrum of questions including the following ongoing research projects:
- Determination of the quantitative genetic, molecular, and genomic basis of blood feeding in W. smithii, the only species of mosquito that bites in one part of its range (southern USA) and never bites in the rest of its range (northern USA & Canada). Identifying the genes responsible for the transition between blood feeding and obligate non-biting is a key step in the eradication of all mosquito-carried diseases world-wide, including malaria and dengue, yellow fever, West-Nile, Zika, and multiple encephalitises. If there is no bite, there is no mosquito disease transmission, period.
- Identification of the shifting patterns of vector-host interaction during rapid climate change. This predictive arm of our research has far-reaching implications for the application of traditional control measures, with special attention focused on those many blood-borne diseases for which there are, as of yet, no effective vaccines or cures.
- Evaluation of lipid and other metabolic pathways known in all animals to be important in energy production, membrane synthesis, nitrogen excretion/conservation, and iron toxicity during the transition from the developmentally dormant state of diapause to the resumption of active development, with an eye towards human disease interventions.
- Genome assembly and comparative genomics: What constitutes a “gene” in a novel organism?
- Comparative transcriptomics, including diapause (insect dormancy) in response to day length, with implications for adult longevity and aging.
- Identifying the evolutionary and genetic relationships between the two great biological time-keeping mechanisms that organize life on Earth as we know it – the daily circadian clock and the seasonal photoperiodic timer. Although the core genes are well defined in the daily clock, not a single gene unique to the seasonal timer has been identified in a natural population of any animal.

- Determination of geographic variation over both latitudinal and altitudinal ranges in the core daily clock genes that control hormonal, metabolic, and behavioral rhythms in all animals, with important applications in agriculture, conservation, and medicine.
- Determination of the relative roles of season length, length of day, and temperature in driving evolution in plant and animal species during rapid climate change. Our lab made the startling discovery that the effects of climate change have penetrated to the level of the gene. This discovery was later confirmed in plants, birds, and mammals.
We currently enjoy collaborations with others within the University of Oregon as well as outside collaborations with Indiana University, University of California Davis, University of Birmingham (UK), University of Notre Dame, Kansas State University, Stowers Institute for Medical Research, Ohio State University, Georgetown University, USDA, University of Florida, Florida State University, and Oregon Health and Science University. We typically have mentored 10-15 students in the lab from undergraduates to post-docs at any given time.




